1. Introduction
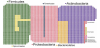
The number of bacteria inhabiting the human body is estimated to be approximately ten times larger than the number of human body cells. It should be mentioned however that this estimate recently has been challenged [1]. The majority of these micro-organisms are found in the gastrointestinal tract, which comprises approximately 1014 bacteria [2-4]. This bacterial population is referred to as the gut microbiota and weighs more than 1.5 kg. Because the human host and the gut microbiota are totally dependent on each other, collectively they are termed a ‘super organism’ [5]. The gut microbiota can be considered an ‘acquired’ organ, because intestinal colonization starts immediately after birth and evolves as we grow. This process could be envisioned as the creation of the (microbial) world in the intestines (Figure 1). The microbiota constitutes an ecologically dynamic community with a number of essential functions for the human host. These include digestion of food components, providing nutrients and vitamins, regulation of intestinal development, maturation and regulation of the immune system, and more, all contributing to overall health [6,7]. In total, 2172 bacterial species have been identified in the human gut (Figure 2) [8], but one individual harbors around 160 different species [9]. Until the 1990s, knowledge of the gut microbiota was limited because conventional culture was the only technique to characterize its composition. However, it is estimated that at least 20 to 60% of the bacteria is uncultivable, resulting in an underestimation of gut microbiota diversity [10]. The introduction of culture-independent molecular and high-throughput approaches enabled phylogenetic investigations and quantification of the bacterial community, thereby refining the original perspective on composition and dynamic changes of the gut microbiota. These techniques moreover advanced our insights into the impact of host and environmental factors on bacterial community structure and dynamics [11].
One of the most exciting developments in recent years in the biomedical sciences has been the understanding that the intestinal microbiota strongly affects human physiology, by elucidating the mechanisms underlying the associations between gut microbiota, health and disease [12-14]. The increase in the number of publications on the human microbiota has been exponential in the last few years and continues to grow (Figure 3). The outburst of discoveries in the microbiome field in the course of the last five to six years has changed our perspective on human biology, both in terms of health and disease.
This manuscript deals with the development of the intestinal microbiota in infancy. It includes the study of the effects of probiotics in early childhood on the composition and functionality of infant microbiota with a focus on the interaction between gut microbiota and development of the neonatal immune system. As search strategy of this review we searched Pubmed (https://www.ncbi.nlm.nih.gov/pubmed/) and Web of Science (http://apps.webofknowledge.com) using the keywords microbiota in combination with development, infant, childhood, as well as immune system and immunity. No restriction on publication year was applied and for review articles the most recent ones were included.
2. Origin and Definitions
The potential association between the composition of human gut microbiota and the development of disease was proposed already by Metchnikoff in the early 20th century. He hypothesized that replacing or diminishing ‘putrefactive’ bacteria in the gut with lactic acid bacteria could normalize bowel health and prolong life [15,16]. Joshua Lederberg, a Nobel prize winning American molecular biologist, known for his work in microbial genetics, emphasized the importance of the intestinal bacterial population and introduced the overarching term ‘microbiome’, ‘to signify the ecological community of commensal, symbiotic, and pathogenic microorganisms that literally share our body space’[17]. Sometimes the term ‘microbiome’ is used interchangeably with the term ‘microbiota’ [18]. The microbiome is the collection of genomes of all microbes that live inside and on the human body (bacteria, but also bacteriophages, fungi, protozoa and viruses). The microbiota is the ecological community of commensal, symbiotic and pathogenic microorganisms that resides in a previously established environment [12]. This microbial community was previously termed gut flora, a term which should be discouraged because it would suggest that bacteria belong to the plant kingdom. Description of the functionality of the gut microbiota can be done by investigating the collective assortment of metabolites present in a stool sample (the fecal metabolome), referred to as ‘metabolomics’; by studying the total transcribed RNA, referred to as ‘metatranscriptomics’, and by focusing on protein levels, called ‘metaproteomics’. ‘Metagenomics’ (also known as environmental genomics or community genomics) is the study of the metagenome, i.e. the collective community of genomes from a particular ecosystem (in this case, the gut). When only genes are assessed, a marker gene study is performed [4,19,20]. It is estimated that the human gut metagenome consists of 9,879,896 genes [21], a dazzling number when compared to the 18,877 human genes [22].
3. Classification and Composition of Gut Microbiota
All life on earth, from microbes to larger organisms, including man, has gone through a similar biochemical evolution. Back in 1977, Carl Woese performed comparative studies of rRNA sequences and underpinned the basis for a universal phylogeny. He outlined a universal sequence-based tree of life, with three phylogenetic domains: Archaea, Bacteria and Eukarya [23]. Nowadays, this three-domain model is grounded by further information on gene sequence and on biochemical correlations [24]. Over the past decades, the rRNA gene sequences have been a major advancement for microbial identification. Most bacterial species isolated from the human gut (93.5%) are members of four bacterial phyla: Firmicutes, Proteobacteria, Actinobacteria, and Bacteroidetes (see also Figure 2) [8]. The Firmicutes are mainly represented by the genera Clostridium, Faecalibacterium, Blautia, Ruminococcus, and Lactobacillus, and the Bacteroidetes including Bacteroides and Prevotella [25,26]. Other phyla such as Proteobacteria (Gammaproteo bacteria with Enterobacteriaceae), Actinobacteria (Bifidobacterium), Fusobacteria and Verrucomicrobia (Akkermansia) are present in lower abundance but also have a significant influence on the total microbial configuration [8,27].
4. Development of the Intestinal Microbiota
From birth onwards, the infant intestine becomes increasingly colonized with a wide variety of microorganisms. This would imply, and that is the general consensus, that during the fetal period the intestine is sterile. Lately this view has been challenged and the concept of a fetal microbiome has been put forward. Is there something like a fetal microbiome, and if so, where does it come from? Sensitive molecular methods would point towards the existence of a placental microbiome, resembling the oral microbiome [28]. Also amniotic fluid and meconium have been shown to contain bacterial DNA [29,30]. Critics state that the placenta microbiome is indistinguishable from contamination during DNA isolation and this issue certainly is not resolved [31].
Whether or not the intestines are sterile at the moment of birth, the gastrointestinal tract will become exposed to bacteria from the environment already during the birth process or immediately after birth. The classical early colonizers of the infant gut are facultative anaerobes of the phylum Proteobacteria, such as Escherichia coli and other Enterobacteriaceae. These organisms pave the way for strictly anaerobic bacteria such as Bifidobacterium, Clostridium, and Bacteroides, and sometimes Ruminococcus, by depleting the initially available oxygen in a matter of days. The rapid colonization by commensal bacteria has great impact for the development of the (mucosal) immune response of the infant. It leads to a balanced development of the various components of the mucosal immune system and induces tolerance to those bacteria. The infant gastrointestinal community is characterized by low stability, limited bacterial richness and great inter-individual variation [32,33]. During the period prior to weaning the microbiome can display large shifts in the abundances of bacterial taxa [34,35]. Subsequently, the intestinal microbiota of the infant slowly develops and matures. Weaning results in a shift towards the bacterial group’s characteristic of the adult microbiota [36]. The microbial community is thought to reach an adult-like composition around three years of age [12,34]. Other literature however indicates that the evolvement into adult-like microbiota may continue beyond that age [37-39]. Which bacteria should be represented (quality as well as quantity) in a ‘healthy’ or ‘normal’ intestinal microbiota has not been clearly defined yet.
Recent literature suggests a broad core microbiota in adults with high variation in species, diversity and genetic functions between individuals, although the main microbial gene functions may be maintained in almost every individual [40-44].
5. Genetic and Environmental Factors Influencing Development of the Intestinal Microbiota
A wide range of factors, genetic and environmental, can influence the diversity and composition of the intestinal microbiota and its establishment (Figure 4). In the first few hours of life, the mother’s vaginal and fecal bacterial populations are usually the most important source of inoculum. Moreover, skin microbiota of parents and siblings, bacteria from breast milk, and bacteria from the environment are sources of strains colonizing the gut in early life [3,43,45]. Mode of delivery, gestational age and feeding mode especially affect the infant’s microbiota composition [46-49]. Besides, prescription of antibiotics in early life and supplementation with prebiotics or probiotics can have a direct and major effect on the development of the intestinal microbiota [47,50,51].
Generally, the mode of delivery is the most influencing factor for microbiota development. Bifidobacterium species, known as beneficial species for the host, dominate the relatively simply composited microbiota of full-term vaginally-delivered infants within the first two weeks of life. Prominent genera are also Lactobacillus, Prevotella, Escherichia and Bacteroides, including a bacterial community that represents the mother’s vaginal and intestinal microbiota [52,53]. Caesarean-delivered infants have a reduced number of Bifidobacterium; Staphylococcus, Corynebacterium, and Propionibacterium are dominating bacterial species in their microbiota which shows less resemblance to their mother’s, as compared to vaginally delivered infants [52,54,55].Another significant factor which influences microbiota composition is feeding type. The microbiota of breast-fed neonates is dominated by the genera Bifidobacterium and Ruminococcus. Formula-fed neonates have a more complex composition compared with breast-fed neonates and harbor a diverse microbiota including Enterobacteriaceae, Enterococcus and Bacteroides, as well as Bifidobacterium and Atopobium [46,56,57].
There is an overwhelming amount of evidence that exposure to antibiotics in early life, which has become common in modern obstetric and neonatal practice, is associated with profound effects on the neonatal gut microbiota composition. In general, antibiotic treatment leads to a decrease in the microbial diversity [58]. Overgrowth of Enterococci and arrested growth of Bifidobacterium in term infants exposed to antibiotics in the first week of life have been described [50,60-62]. Antibiotic exposure at the beginning of life can modify the course of bacterial infections and is associated with development of atopic disease [63-66] and with other immune mediated diseases [67,68] as well as obesity [69].
Information on the role of genetic factors which affect the composition of the intestinal microbiome comes from studies of monozygotic and dizygotic twins and studies in humans where specific microbiota profiles are associated with subjects that have mutations at specific genetic loci. The data show that the microbiome is shared among family members, because individuals from the same family (twin pairs, or twins and their mother) have a more similar microbiota than unrelated individuals [70-72]. Despite the wide variation of shared microbial genes among individuals, an extensive, identifiable human 'core microbiome' at the gene level could be identified and deviations from this core were associated with different physiological states [42]. An increasing number of genes/molecules (such as the genes of the class 2 major histocompatibility complex (MHC)) is found to be associated with specific microbial compositions and detection and monitoring of the intestinal lumen by the mucosal immune system of the gut [73].
Microbial colonization of the intestine plays an important role in the postnatal development of the gastrointestinal system [74], thereby influencing host metabolism and disease development [18,20,75-77]. Bacteria form a complex ecological community that can modulate the expression of genes involved in several important intestinal functions. The community influences normal physiology and susceptibility to disease through collective metabolic activities and interactions, including nutrient absorption, mucosal barrier strengthening, xenobiotic metabolism, angiogenesis and postnatal intestinal development. Other functions of gut bacteria are the involvement in the metabolism of otherwise indigestible carbohydrates and the production of essential metabolites such as vitamin K [78-82].
6. Impact of Gut Microbiota on Development of the Immune System
The development of the immune system starts already during the fetal period. After 8 weeks of gestation the first B lymphocytes can be detected in the fetal liver. Progenitors of T lymphocytes start migrating to the thymus anlage by weeks 8 to 9. B lymphocytes are detectable in fetal blood circulation by 12 weeks of gestation, T lymphocytes 3-4 weeks later [83]. The functional immune repertoire further develops during fetal life, but is not challenged because of the sterile environment [84-87].
The function of the mucosal immune system is to prevent pathogenic micro-organisms from the environment (in the context of the present discussion: from the gut) to invade the body. As outlined above, immediately after birth the gastrointestinal tract becomes colonized with bacteria and other micro-organisms.
The microbiota that colonizes the neonatal gastrointestinal tract is of utmost importance for growth and maturation of the various cellular components of the intestines itself, but certainly also for the regulated development of both the mucosal immune system of the gut as well as the systemic immune system [88]. This is demonstrated by (severe) deficiency of various types of immune cell types and organized lymphoid tissue in germ-free animals: secondary lymphoid organs such as mesenteric lymph nodes and Peyer’s patches are smaller and fewer in number and have a poorly organized structure. T and Blymphocytes are reduced in numbers [88]. The gut microbiota also modulates the development and functionality of the innate immune system lining the gastrointestinal tract [89-91].
It would be an immense, if not impossible task, for the immune system to eliminate all micro-organisms from the gut. The immune system therefore is programmed and matures in such a way that resident, non-pathogenic micro-organisms are tolerated and compartmentalized [92]. Induction of tolerance to resident microbiota is induced immediately after birth and involves Toll-like receptor (TLR) signaling via secretory leukocyte protease inhibitor proteins [93,94].
A prominent deficiency of germ-free mice is the lack of expansion of CD4+ regulatory T lymphocytes (Tregs) [95]. This deficiency can be completely restored by the treatment of germ-free mice with polysaccharide A from the capsule of Bacteroides fragilis (see also Figure 5, panel 1) [96-98]. This process is mainly mediated by the pattern recognition receptors (PRRs) of epithelial cells, such as TLRs or Nod-like receptors, receptors which are able to recognize molecules (such as polysaccharide A) produced by intestinal microbes. These mechanisms allow discriminating between beneficial and pathogenic bacteria and increasing the number of immune cells or PRRs [99-101].
B. fragilis is not the only bacterial species that can induce mucosal Tregs; also Roseburia and various Clostridia species as well as others have this capacity [102,103].
In this case the mechanism is totally different. Human colonic bacteria can ferment resistant starch and non-starch polysaccharides (which are major components of dietary fiber) to short-chain fatty acids (SCFAs). Among the SCFAs, butyrate, that has multiple functions including anti-inflammatory properties, is important for a healthy gut physiology [81,82]. The SCFAs, in particular butyrate and acetate, can directly promote Treg differentiation in the colon of mice (see also Figure 5, panel 3) [104-106]. The butyrate acts as a transcription factor via histone deacetylase inhibition and also via a G protein-coupled receptor signaling mechanisms [106].
Another example of a commensal bacterium with a direct immunoregulatory function is the segmented filamentous bacterium SFB (see Figure 5, panel 2). SFB belongs to a class of anaerobic and clostridia related spore-forming bacteria of the gastrointestinal tract [107]. SFB is closely associated with gut epithelial cells and stimulates these cells to release serum amyloid A1. Colonization of germ-free animals with SFB stimulates maturation of the gut mucosal lymphoid tissue, promotes mucosal IgA production, and stimulates the differentiation of T-helper 17 cells [108].
Akkermansia muciniphila (the single representative of the phylum Verrucomicrobia in the human gut; see Figure 2) is associated with protection against several inflammatory diseases [109,110]. Both the expression of type IV pilias well as production of SCFAs have been proposed as mechanisms of action [111,112]. In patients with Crohn’s disease a lower abundance of Faecalibacterium prausnitzii is found [113]. An anti-inflammatory protein from F. prausnitzii has been shown to inhibit NF-κB in intestinal epithelial cells and (thus) prevents colitis in an animal model [114,115].
All these examples underscore the importance of gut microbiota on development and function of the (mucosal) immune system. The most compelling evidence for modulation of the immune system by gut microbiota comes from the response to checkpoint inhibition therapy and adoptive cell therapy for cancer [116-122]. Elucidation of the underlying mechanisms and identification of the key microbial species will be the next steps.
7. Modulation of Intestinal Microbiota by Probiotics and Prebiotics
The gut microbiota plays a crucial role in the physiological development of both the intestinal as well as the immune system, and therefore disturbances in gut microbiota can be causally related to inflammatory and immune mediated diseases. Indeed, the development of allergic diseases has been linked to an altered gut microbiota composition, reduced microbial exposure and reduced bacterial diversity in childhood [123-125]. This hypothesis has been modified into the ‘gut microbial deprivation hypothesis’, with its emphasis on alterations of original gut microbiota during infancy [126-130]. Whatever the cause, an aberrant microbial colonization pattern or a distortion of the microbial ecology early in life might predispose the infant to immune mediated diseases such as T-helper 2 (Th2)diseases like allergy, wheezing and asthma, or auto-inflammatory T-helper 1 (Th1) diseases, like inflammatory bowel disease, diabetes and obesity [75,125,131-135]. In this context, the window of opportunity for influencing the composition of the intestinal microbiota and possibly modulation ofthe development of (later) allergic or autoimmune diseases, would be the neonatal period.
There is increasing evidence that modulation of the infant microbiota can restore the ecological balance [137-139]. This modulation is mainly done by supplementation of pre- and probiotics. A prebiotic is defined as ‘a selectively fermented ingredient that results in specific changes, in the composition and/or activity of the gastrointestinal microbiota, thus conferring benefit(s) upon host health’ [140].The term ‘probiotic’ is used for microorganisms with a health benefit beyond basic nutrition. Probiotics are defined as live microorganisms that, when administered in adequate amounts, confer a health benefit on the host [141]. The most widely used probiotic bacteria belong to either the Lactobacillus or the Bifidobacterium genera. Many studies have investigated the potential of probiotics, both for primary prevention as well as treatment for infectious and/ or immune-mediated diseases. The Cochrane database of systematic reviews list a number of infectious and immune-mediated diseases for which probiotics have shown to have a beneficial effect. These include prevention of Clostridium difficile-associated diarrhea, both in adults as well as in children [142], pediatric antibiotic-associated diarrhea [143], the prevention of acute upper respiratory tract infections [144] and of ventilator associated pneumonia [145], the severity of hepatic encephalopathy [146], the treatment and prevention of pouchitis after ileal pouch-anal anastomosis for chronic ulcerative colitis [147], and prevention of necrotizing enterocolitisin preterm infants [148]. Results obtained with probiotics for primary prevention or treatment of allergic diseases are variable [148-154], which may be related to the strain-specific effects of probiotic bacteria and/ or considerable variations in study design, doses administered and duration of the intervention. In a recent systematic review of diet during pregnancy and infancy and the risk of allergic or autoimmune disease development, a relationship was found between maternal diet during pregnancy and lactation, and eczema or allergic sensitization to food during childhood. A positive health effect was also found for probiotics and supplements of fish oil [155].
Modulation of the (composition and functionality) infant gut microbiota can also be executed by prebiotics. This topic has recently been dealt with in a series of excellent reviews [156-158]. Prebiotics, either alone or in combination with probiotics (the combination of the two is called synbiotics) thus help to develop a healthy gut microbiome. Breast milk, containing human milk oligosaccharides, is a natural source of prebiotics and provides the baby with optimal active ingredients for outgrowth of especially bifidobacteria [159].
One of the potential mechanisms by which probiotic bacteria regulate the development of the mucosal immune system is by stimulation of the differentiation of naive T cells towards Th1 or Treg cells, thereby shifting the balance between Th1 and Th2 cells. There are indications that probiotic intervention, via induction of butyrate production, actually could work this way [160]. Future studies should address host and microbiota interactions and identify optimal timing and duration of pre- and/or probiotic supplementation as strategy for prevention of immune-mediated diseases.
8. Summary and Conclusion
To summarize: the human infant develops an initial microbiota as it becomes colonized with a wide variety of microorganisms during or directly after birth. The neonatal period comprises a critical period for its development, when early programming occurs and the immune system matures. The long-term composition and functionality of the newborn’s gut microbiome is programmed during this period, thereby laying the foundation for future health and influencing the risk of developing disease later in life [161,162]. The development of the gut microbiota continues during the first years of life and its composition is considered to resemble the adult gastrointestinal tract when the child is at the age of 3 years [38,163]. Modulation of the infant microbiota can restore the ecological balance of the microbiota and thus contribute to an optimal development of the immune system, enabling adequate defense against infectious diseases, while avoiding allergic and autoimmune diseases.
Competing Interests
The authors declare that they have no competing interests.
Author Contributions
NBR and GTR are responsible for conception and design of the review. All authors contributed to analysis of the literature, writing, and critical reflecting of the manuscript. All authors have seen and approved the final version of the manuscript.