1. Introduction
Mesenchymal stem cells (MSCs), which are capable of self-renewal and differentiation into various tissues, have attracted attention for use in regenerative medicine [1-4]. Many studies have demonstrated that MSCs can differentiate into osteoblasts, chondrocytes, adipocytes, muscle cells and even neuronal cells, and show respective functions in vivo after transplantation [1-6]. One of the approaches to utilize MSCs in regenerative medicine is transplantation of fully differentiated cells which are differentiated from MSCs in vitro [5,6]. The traditional method to induce the specific differentiation of stem cells in vitro is based on the treatment of chemicals such as several growth factor mixtures that lead stem cells to a specific lineage [5-8]. However, recent studies suggest that mechanical factors, such as rigidity or surface topography of the cell culture substrates, can also affect the stem cell fates [9-13]. For example, the surface topography in groove and ridge patterns, which have micron-scale widths and nanoscale depths, induced osteogenesis of the MSCs [12], whereas nanotopography in groove and ridge patterns (300 or 500 nm) induced neuronal differentiation of MSCs [13]. In addition, a recent report showed that MSCs can differentiate effectively on a substrate with disordered asymmetric post patterns, in contrast to the substrate with ordered post patterns, and demonstrated that symmetry or regularity of the surface topography is also important for the decision of the stem cell fates10. Another study demonstrated that soft matrices (0.1 ~ 1 KPa) induced neurogenic progenitor cells, while stiffer matrices (8 ~ 17 KPa) and rigid matrices (25 ~ 40 KPa) induced myogenic and osteogenic progenitor cells, respectively [11].
However, it has not been investigated whether cells can distinguish and respond to each mechanical cue when diverse combinations of various mechanical stimuli are given simultaneously. In addition, it has not been studied whether a specific mechanical cue can predominantly affect cellular behaviors among various mechanical cues. For example, even though it was known that the rigid substrate tends to direct osteogenesis of MSCs [11] and nanogroove topography induces neural differentiation of MSCs [13], there is no information about the differences between the cellular responses on the nanogroove patterns made of rigid materials and soft materials. However, this kind of research provides useful information to develop and optimize the cell culture substrates for specific stem cell differentiation.
Furthermore, it is also necessary to investigate the alteration of cytoskeletal systems while diverse combinations of various mechanical stimuli are given simultaneously to cells. Recent studies showed that cells respond to mechanical stimuli through the cytoskeletal system, which discerns the physical changes of microenvironments around the cells and passes the information on to down-stream of the signal pathway to regulate cellular shapes and differentiation14-17. These results were also supported by several recent studies that showed that disrupting the cytoskeletal systems inhibited mechanical-cuemediated morphology change and fate decision in stem cells11, 17. However, it is still elusive how cytoskeletal systems react to the diverse mechanical cues at the same time, and how they integrate the several mechanical signals.
Thus, we were motivated to investigate whether two different mechanical cues can simultaneously influence the cytoskeletons and cellular shapes. In this study, we examined the cellular shapes and the structures of actin cytoskeletons in MSCs on different elastic matrices with the same surface topography in various dimensions.
2. Materials & Methods
2.1 Cell Culture
Human MSCs were provided from PromoCell (Heidelberg, Germany) and maintained in low-glucose Dulbecco's Modified Eagle's Medium (DMEM with 1 g/L glucose) containing 10% fetal bovine serum (FBS), 100 units/ml penicillin, and 100 μg/ml streptomycin [2,3]. The cells were incubated at 37°C and 5% fully humidified co2. Cells in passage 6 were used for all experiments in this study.
2.2 Substrate preparation
Substrates for cell culture were fabricated on two different materials, polydimethylsiloxane (PDMS) and SU-8. For patterning structures on PDMS substrates, PDMS were cast on a master mold which was photo-lithographically fabricated in SU-8 on a silicon wafer, or master patterns in etched silicon surfaces after e-beam lithography. Micropatterned SU-8 substrates were fabricated on tissue culture plates (TCP) by soft lithography18 using a PDMS replica mold. In detail, PDMS replica molds were placed on SU-8-coated TCP surfaces and exposed to UV-rays for 10 min., and the PDMS molds were removed and cured for 20 min. For nanoscale patterns, since soft lithography using PDMS molds cannot guarantee accurate features, a hard polyurethane acrylated polymer mold coated with an anti-adhesion PDMS layer19 was used for the nanopattern transfer process. The rest of the process is the same as the microscale SU-8 molding described above. All the substrates were prepared for cell culture by coating with 10 μg/ml fibronectin for 2 h in a 5 % co2 cell culture incubator. PDMS substrates were treated with oxygen plasma before the coating.
2.3 Cell staining
After the cells were cultured on various substrates, they were fixed in 3.7 % formaldehyde for 1 h, rinsed in phosphate buffered saline (PBS), and permeabilized by treating with 0.1 % tritonX-100 for 15 min. Then, actin cytoskeletons were stained with rhodamine-conjugated phalloidin, and the nuclei were stained with 4'-6-diamidino-2- phynylindole (DAPI).
3. Results and Discussion
3.1 Various substrates for cell culture
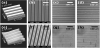
To compare the cellular shapes and the organization of the actin cytoskeletons in MSCs on different elastic matrices with the same surface topography, we selected two polymers: polydimethylsiloxane (PDMS) as a soft polymer and SU8 as a rigid polymer. PDMS is a silicon elastomer whose Young’s modulus is around 1 MPa, and SU8 is a highly rigid epoxy-based polymer whose Young’s modulus is 2~5 GPa. The surface topographies were created by replicating micro- or nano-patterns from a master mold on each polymer by soft lithography or nanopattern transfer technique [19], which are described the materials and methods section. Figure 1 show all the patterned PDMS and SU8 substrates used as matrices for culture of MSCs throughout this study. The dimensions of the groove and ridge patterns are 0.5 × 0.5 × 0.5 μm (Figures 1a and 1e), 1 × 1 × 1 μm (Figures 1b, and f), 20 × 20 × 10 μm (Figures 1c and 1g), and 50 × 50 × 10 μm (Figures 1d, and 1h) (groove width × spacing width × depth). As shown in Figure 1, micron-scale patterns made of both PDMS and SU8 presented high accuracy in their structures (Figures 1 b, c, f, and g), but the nanoscale pattern (0.5 μm groove width × 0.5 μm spacing width × 0.5 μm depth) in PDMS (Figure 1e) displayed less structural accuracy than the same pattern in SU8. Thus, in the following experiments, the effects of PDMS and SU8 patterns on MSCs were compared to one another on only the micron-scale patterns. Results from the culture on nanopatterned surfaces were considered as reference data.
3.2 Morphological change of MSCs grown on various substrates
To evaluate the morphological changes of MSCs on the micro and nanotopography made of two different polymers, cells were cultured on each substrate, and F-actin was stained with rhodamineconjugated phalloidin to observe the boundary shape of the cells using microscopy (Figure 2). As shown in Figures 2(a-c), the shape of MSCs were not significantly changed in the non-patterned PDMS and SU8 substrates, compared to MSCs on tissue culture plates. The cells were stretched well on all the substrates, but their orientations were irregular. In a previous report, it was suggested that MSCs showed significant changes in their morphology within a few hours when they were grown on different elastic matrices with flat surfaces11. In our study, the elasticity of the materials was between about 1 MPs and 2 GPa, whereas the stiffness of the matrices used in the previous paper was between 0.1 KPa and 40 KPa. We assumed that the cells could not sense the difference in stiffness in this range, and this might also be a reason for why changes in cell shape were not clearly observed on the different flat substrates in our results. To elucidate the correct reason for this phenomenon, further studies are needed for the identification of the threshold values of the stiffness ranges sensed by MSCs on different elastic matrices.
When MSCs were cultured on the patterned substrates with the dimensions of 0.5, 1, 20, and 50 μm (groove and spacing width, Figures 2(d-k), the cells oriented along the direction of patterns such that the longest axes of the cells were aligned in parallel to the grooves and ridges. In addition, the cells on the PDMS groove and ridge patterns were elongated more than those on SU8 substrates with the same pattern of structures. On the PDMS substrate with 1-μm groove patterns, it was commonly observed that cells were extremely elongated more than 1 mm. In addition, some parts of the cells on that substrate showed morphological similarity to neuronal cells composed of a single cell body and highly branched neuritis, while this was not observed on the SU8 1-μm groove patterns. This result indicates that cells can sense the surface topography and matrix elasticity simultaneously. Furthermore, it is very interesting that the differences in the cell elongation and shape change on the PDMS and the SU8 substrates were observed only on the pattered substrates (especially on the 1-μm groove patterns), while no difference was observed on flat surfaces. Through this result, we propose that there are some interactions between sensing the matrix elasticity and sensing the surface topography in the cells.
3.3 Actin-Alignment of MSCs grown on various substrates
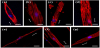
To corroborate the results (Figure 2), high-magnification images (×800) of actin cytoskeletons were taken using a confocal laser scanning microscope (CLSM). Figure 3 shows some representative images of the cells cultured on the same substrates previously used for fluorescent microscopic imaging. Actin cytoskeletons were stained with rhodamine-conjugated phalloidin, and the nuclei were countstained with DAPI. In the cells on the SU8 substrates with 0.5-μm groove patterns, actins were strongly expressed only at the edge of the cells (Figure 3a). On the SU8 substrates with the 1-μm groove patterns, actin fibers aligned in parallel to the patterns were clearly observed in both the middle and the edge of the cells (Figure 3b). In the case of the 20-μm groove made of SU-8, even though the cell bodies were aligned along the patterns, most of the actin fibers inside the cells were biased by about 60 degrees (Figure 3c). However, actins on the 50-μm patterns were not organized in a specific orientation; these actins were biased and formed meshwork. On the patterned PDMS substrates, strong actin fibers were not observed as much as on the patterned SU8 surfaces (Figures 3e-g). It was observed that the filament-like actins were aligned along the pattern, and the actins formed meshwork on all the patterned PDMS substrates.
Subsequently, different cell shape parameters, such as the degree of elongation, distinct form, and alignment of actin cytoskeleton, were observed more significantly on substrates with different stiffnesses compared to flat substrates. Unfortunately, sufficient data were not accumulated to explain these phenomena. Further investigation is necessary to understand the mechanism of the substrate stiffness effect on the dynamic rearrangement of cytoskeletons in MSCs.
4. Conclusion
We have demonstrated that the matrix elasticity and the surface topography simultaneously affect cell morphology and cytoskeleton rearrangement. Furthermore, our results reveal that the matrix elasticity (1 MPa and 2 GPa) has a greater effect on changes of MSC shapes and cytoskeletons with surface topography than without surface patterns. Therefore, we propose that there are some interconnections between sensing the matrix elasticity and sensing the surface topography in the cells. In conclusion, the combination of the matrix elasticity and the surface topography are considered to apply matrix mechanical cues for specific differentiation of stem cells in regenerative medicine.
Competing Interests
The authors have declared that no competing interests exist.
Author Contributions
SH Kim performed the experiment and data analysis, G Lee and S Park did analysis and wrote the manuscript. The funding for the research was acquired by S Park.