1. Introduction
Platinum compounds, the DNA-targeting drugs, are effective and have been widely used in clinical settings. However, acquired resistance to platinum-chemotherapy is a major treatment obstacle affecting life-quality of patients and therapeutic outcomes, often leading to treatment failure [1,2]. Studies show that cells treated with genotoxic agents swiftly respond by activating DNA-damage checkpoint response. Two primary pathways are initiated in response to DNA damage. One is mediated by the ATM-Chk2 axis, the other by the ATR-Chk1 axis. The ATM-Chk2 pathway responds primarily to DNA double-strand breaks, whereas the ATR-Chk1 pathway mainly responds to replication-associated DNA lesions [3,4].
Drug-resistance is multifactorial in nature. Its mechanisms involve a complex network of cellular pathways and molecular changes with numerous cross-interactions at different stages. Resistance to platinum-compounds may occur due to alterations of drug influx and efflux; altered activation or metabolism; modified drug-induced damage or drug-targets; and/or evasion of apoptosis [5,6]. To date, however, no successful chemosensitizers or diagnostic/prognostic assays for the prediction of therapy response have been developed.
Transcription factor p53 plays a key role in the DNA damage response to genotoxic stress. Wild type p53 protein has a role in the inhibition of DNA synthesis following DNA damage, suggesting a mechanism for how the loss of wild-type p53 may contribute to tumorigenesis [7,8]. Mutations of p53 have been associated with resistance to platinum-based chemotherapy and shortened survival in ovarian cancer [9].
Checkpoint kinase 2 (Chk2) resides at the heart of the DNA damage/repair pathway and is responsible for the maintenance of mammalian genomic integrity. Studies suggest that Chk2 inhibition in combination with genotoxic agents might have therapeutic value [10-12]. Inhibition of Chk2 expression reduces DNA-damage-induced cell cycle checkpoints and enhances apoptosis in p53-defective HEK- 293 cells [11]. Molecular or genetic targeting of Chk2 prevents the release of survivin from mitochondria and enhances DNA-damage -induced tumor cell apoptosis, thus inhibiting in vivo growth of resistant tumors, providing a rational approach for treatment [13,14].
2. NER Pathway is Associated with Platinum Resistance Phenotype
Nucleotide excision repair (NER) is the critical DNA repair pathway.
It appears to be the major mechanism for removal of platinuminduced DNA adducts, resulting in resistance to platinum drug therapy. NER enzyme-complexes remove bulky, transcription blocking lesions caused by endogenous and environmental insults to DNA, including platinum-induced adducts [15-17]. NER-defective cells are hypersensitive to platinum drugs, and enhanced DNA repair has been implicated in the cisplatin-resistant phenotype [18-20]. The major steps in the NER process are damage recognition, dual-incision of damaged DNA (on both 5' and 3' sides of the lesion), removal of incised nucleotides and deoxyribose, and gap fill-in synthesis [21-26].
3. ERCC1 Expression Predicts Chemosensitivity
ERCC1 is an essential component of the NER pathway, which is the only known mechanism for the removal of intrastrand/interstrand bulky DNA-adducts. Studies indicate that high levels of ERCC1 expression reflect elevated DNA repair capacity and are associated with clinical resistance to platinum-based chemotherapy [18,19,27].
In vitro studies suggest that ERCC1 expression in cisplatinhypersensitive, repair-deficient cells is 50- to 30-fold lower than in platinum-resistant cells [28]. Overexpression of ERCC1 and other NER genes is associated with increased DNA repair activity and clinical resistance to platinum treatment [29,30]. Data from in vitro systems have shown that suppressed ERCC1 expression by siRNA enhances or restores sensitivity of human cancer cells to cisplatin [31]. Other studies also reported that enhanced DNA repair of platinum-DNA adducts or removal of cisplatin-induced interstrand and intrastrand crosslinks has been observed in cisplatin-resistant cell models, and DNA repair inhibitors in some models can potentiate cisplatin cytotoxicity [32-34]. These findings have significant therapeutic implications.
4. Functional Cis-Elements, AP1 and MZF1, in the ERCC1 Promoter Region Responding to Cisplatin Stimulation
In order to control ERCC1 expression, we performed functional analysis of ERCC1 promoter region by Electrophoresis Mobility Shift Assay and revealed two cis-elements AP1 and MZF1 [35]. In response to cisplatin stimulation, the AP1 site and MZF1 site formed DNAprotein complexes. Furthermore, after cisplatin treatment, binding activities of AP1 increased and binding activities of MZF1 decreased during time course. This suggests that AP1 plays a role as an activator and MZF1 as a repressor. We also measured mRNA expression of AP1 and MZF1 affected by cisplatin and observed that both c-jun and c-fos mRNA levels were increased after cisplatin exposure. In contrast, MZF1 mRNA decreased nearly 75% at 48 hours after cisplatin treatment (data not shown) [34,35].
Put together, our data suggest that within the ERCC1 promoter, the region of -220 to -110 is essential to constitutive ERCC1 expression. And a more forward upstream region containing activator AP1 and repressor MZF1 binding sites is responsible for cisplatin-induced ERCC1 upregulation. In other words, AP1 and MZF1 are the ciselements reacting to cisplatin stimulation. In response to cisplatin treatment, decreased MZF1 and increased AP1 binding activities appear to be the leading mechanism of up-regulation of ERCC1 expression [34,35].
5. ERCC1 SNP Serves as a Prognostic Marker
In 1997, our group discovered a significant single nucleotide polymorphism (SNP) in the ERCC1 gene (GenBank Acc # AF001925). We identified a C to T change at codon 118 in Exon 4 of ERCC1 gene. This change converts a common codon usage to an infrequent codon usage, reducing frequency of use 2-fold. We hypothesized that this SNP would be associated with reduced ERCC1 translation and improved response to platinum chemotherapy [36].
The ERCC1 SNP has been shown by other researchers as an important biomarker associated with platinum sensitivity and predicts better overall survival of patients with several cancers treated with platinum-combination therapy. Specifically, following studies in advanced non-small-cell lung cancer patients, Isla and colleagues concluded that patients homozygous for the ERCC1 118 C allele demonstrated a significantly better survival. They suggest that ERCC1 SNP assessment could be an important component of tailored chemotherapy trials [37]; Ryu et at indicated that median survival time in patients showing C/C genotype of ERCC1 polymorphism was 486 days, which was significantly different from the 281 days of patients with the variant genotype (T/T or C/T; P = 0.0058) [38]. Smith et al. suggested that the C/C genotype at codon 118 may benefit from the combination of platinum and paclitaxel in ovarian cancer patients [39].
6. Wild-Type p53, a Monitor of DNA Damage, Plays a Critical Role in Chemotherapy
In 1994, Lowe and colleagues studied p53 status and the efficacy of chemotherapy in vivo. They found that p53 mutations were detected in resistant or relapse tumor-bearing animals. In response to radiotherapy/chemotherapy, MT-p53 mice responded poorly to the initial treatments, comparing to WT-p53 group, indicating that p53 mutation is associated with treatment resistance and tumor relapse [40]. These findings suggest a basis for the association between p53 mutation and drug resistance/poor prognosis; and between p53 mutation and tumor relapse in patients during chemotherapy.
7. Platinum Drugs Induce p53 Phosphorylation, which Modulates Chk2 Activation
In an investigation of cisplatin-induced molecular signature in cisplatin-sensitive ovarian cancer A2780 cells, we found that several kinases of the DNA-damage repair pathway were activated. One hour after drug exposure we observed phosphorylations of p53 at serine 15 and serine 20, and Chk2 at threonine 68, and increased proteins of ATM, p53, p48 and p21. Of note, cisplatin induced p53 phosphorylation was 12-h earlier than Chk2 phosphorylation, which suggests that Chk2 is activated and regulated by p53 in a wild-type p53 cell model [12]. In another investigation of dicycloplatin-activated DNA-damage response pathway in the same cells, two major factors - p53 and Chk2 - were activated in a manner similar to cisplatininduction, suggesting that the molecular mechanism of dicycloplatin anti-cancer activity is similar to cisplatin [41].
8. Overexpression of p53 Increases Chk2 Phosphorylation in Wild-Type p53 Cells but Not in p53-Null Cells
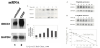
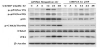
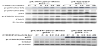
To investigate our hypothesis that only wild-type p53 phenotype possesses the p53 function, cDNA-transfection was performed in both wild-type p53 A2780 and p53-null SKOV3 cells. Overexpressed p53 gene increased cisplatin-induced Chk2 phosphorylation in the wildtype p53 cell model. It doubled the amount of Chk2 phosphorylation 48-h after drug treatment (Figure 3A). In contrast, western analysis showed no effect on Chk2 phosphorylation by cDNA transfection in p53-null SKOV3 cells (Figure 3B). In other words, transfection of p53 in MT-p53 cells failed to alter Chk2 activation [12].
9. Inhibition of p53 by Specific siRNA Inhibits Chk2 Phosphorylation
To confirm the above observations, we performed p53 knock-off siRNA assays and measured Chk2 expression. As shown in Figure 4, in cells not treated with cisplatin, the siRNA to human p53 produced a decrease of phosphorylated Chk2, compared to the nonspecific siRNAtreated control. This decreased level may reflect a constitutive level of activated Chk2 that is normally regulated by p53. Cells transfected by specific siRNA to p53 and treated with cisplatin resulted in a great reduction of phosphorylated Chk2 at Thr-68, suggesting that p53 modulates 68-threonine phosphorylation of Chk2 [12].
These results indicate that in specific conditions Chk2 activation is regulated by p53 in response to cisplatin treatment in wild-type p53 cells but not in p53-deficient cells. Cells without wild- type p53 could survive, conceivably, via an alternative pathway in response to cisplatin treatment. Therefore, we strongly suggest that any research involving the p53 gene should determine its mutational status due to functional differentiation between wt-p53 and p53-mutant.
Competing Interests
The author declare that there is no competing interests regarding the publication of this article.
Acknowledgments
We acknowledge the Molecular Medicine Core Facility, Mary Babb Randolph Cancer Center, West Virginia University and Sopo-Xingda Pharmaceutical, Inc., Beijing, P.R. China for supporting our studies. Special thanks to Michael D. Mueller for editorial assistance.